IMGT Repertoire (IG and TR)
Locus representation: goat (Capra hircus) IGL
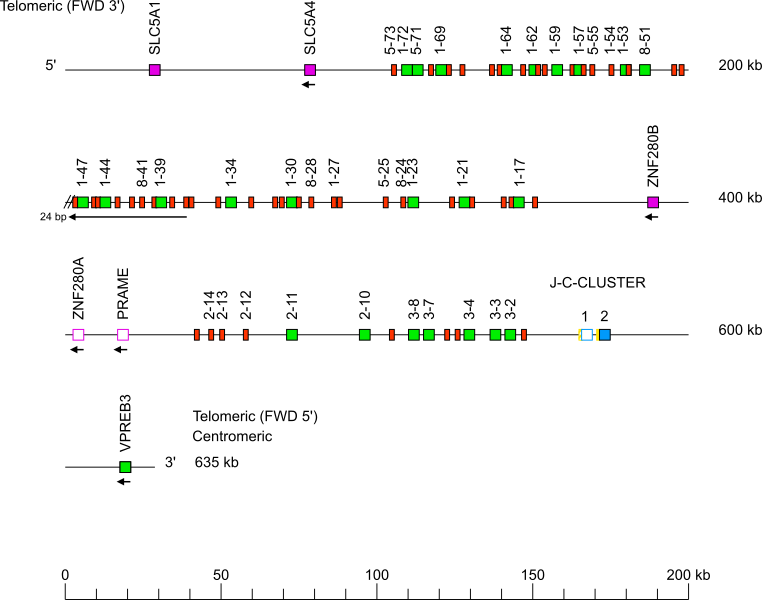
Goat (Capra hircus) IGL locus on chromosome 17
The orientation of the goat (Capra hircus) IGL locus on the chromosome 17 is reverse (REV).

Legend:
Colors are according to IMGT color menu for genes.
The boxes representing the genes are not to scale. Exons are not shown.
A double slash // indicates a gap in genome assembly. Distances in bp (base pair) or in kb (kilobase) associated with a double slash are taken
into account in the length of the lines and included in the numbers displayed at the right end of the lines.
A dotted line ... indicates the distance in kb (kilobase) between the locus and its bornes (5' borne and the most 5' gene in the locus and/or 3'
borne and the most 3' gene in the locus) on the top and bottom lines, respectively. These distances are not represented at scale and/or are not
included in IMGT-LOCUS-UNIT and may not be included in the numbers displayed at the right ends of these two lines
.
IGL gene names are according to the IMGT nomenclature [2].
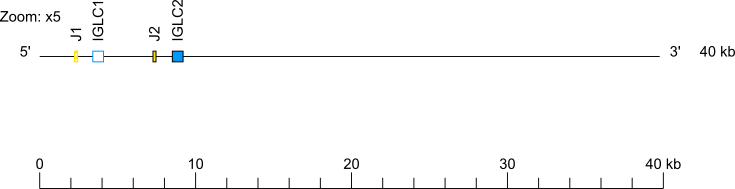
The Capra hircus IGL J-C-CLUSTER comprises 2 cassettes indicated by the numbers 1 and 2 (IGLJ1-IGLC1 and IGLJ2-IGLC2, respectively).
Single arrows show genes whose polarity is opposite to that of the J-C-CLUSTER.
There are two gaps in the locus, one at each end of the inversion, the one in 3', localized in the 5'UTR of IGLV1-37 (complement(236641..236646)) is not represented.
Gene names of IGLV pseudogenes with frameshift(s) in the V-REGION are not displayed due to a lack of space in the locus map at the IMGT standardized scale (line = 200 kb). For all gene names, see Zoom x2.
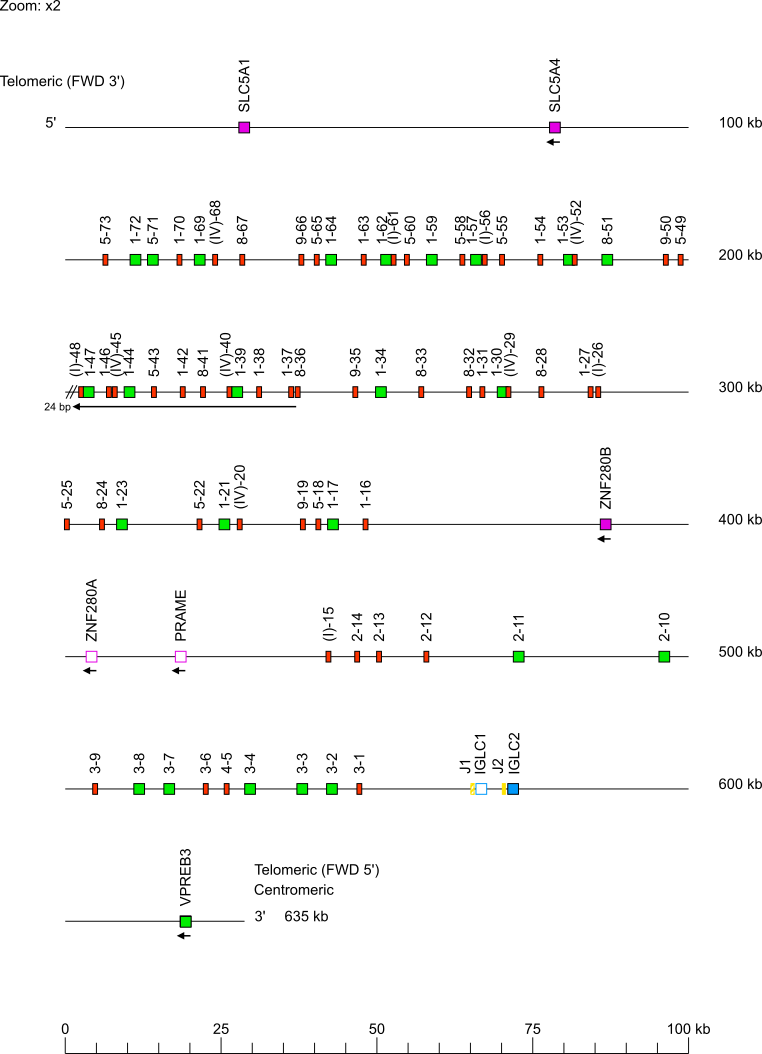
Zoom for IGL V-J-C-CLUSTER

Zoom for IGL J-C-CLUSTER
- [1] Schwartz J.C. et al. Immunogenetics 2018 May;70(5):317-326. doi: 10.1007/s00251-017-1033-3. Epub 2017 Oct 23. Free PMC Article. PMID:29063126
- [2] Lefranc M-P. Front Immunol. 2014 Feb 05;5:22. doi: 10.3389/fimmu.2014.00022. Open access. PMID:24600447
- Created:
- 27/06/2019
- Last updated:
- 22/09/2023
- Authors:
- Morgane Bertignac
