IMGT Repertoire (IG and TR)
Locus representation: bottlenose dolphin (Tursiops truncatus) TRG
IMGT/LIGM-DB: IMGT000015 (56058 bp), bottlenose dolphin (Tursiops truncatus) TRG locus.
Bottlenose dolphin (Tursiops truncatus) TRG locus
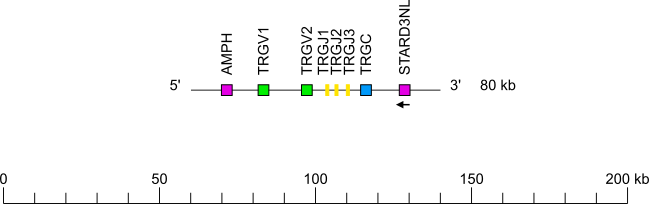
Legend:
Colors are according to IMGT color menu for genes.
The boxes representing the genes are not to scale. Exons are not shown.
A double slash // indicates a gap in genome assembly. Distances in bp (base pair) or in kb (kilobase) associated with a double slash are taken
into account in the length of the lines and included in the numbers displayed at the right end of the lines.
A dotted line ... indicates the distance in kb (kilobase) between the locus and its bornes (5' borne and the most 5' gene in the locus and/or 3'
borne and the most 3' gene in the locus) on the top and bottom lines, respectively. These distances are not represented at scale and/or are not
included in IMGT-LOCUS-UNIT and may not be included in the numbers displayed at the right ends of these two lines
.
TRG gene names according to the IMGT nomenclature [1].
Single arrows show genes whose polarity is opposite to that of the J-C-CLUSTER comprising TRGJ (J1 to J3)-TRGC.
The bottlenose dolphin (Tursiops truncatus) Locus representation is Fig. 1A modified from:
Linguiti G, Antonacci R, Tasco G, Grande F, Casadio R, Massari S, Castelli V, Consiglio A, Lefranc M-P, Ciccarese S.
Genomic and expression analyses of Tursiops truncatus T cell receptor gamma (TRG) and alpha/delta (TRA/TRD) loci reveal a similar basic public γδ repertoire in dolphin and human.
BMC Genomics. 2016 Aug 15;17(1):634. doi: 10.1186/s12864-016-2841-9. Free PMC Article. PMID: 27528257
The original figure was kindly provided by Salvatrice Ciccarese.
- [1] Lefranc M-P. Front Immunol. 2014 Feb 05;5:22. doi: 10.3389/fimmu.2014.00022. Open access.. PMID:24600447
- Created:
- 23/11/2016
- Last updated:
- 22/09/2023
- Authors:
- Saida Hadi-Saljoqi, Perrine Pégorier and Marie-Paule Lefranc
