IMGT Repertoire (IG and TR)
Locus representation: goat (Capra hircus) IGL locus on chromosome 17
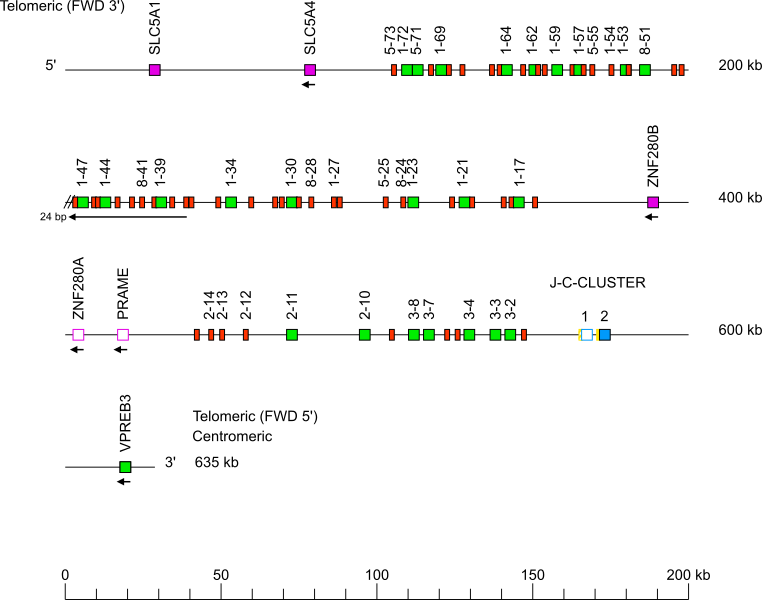
Goat (Capra hircus) IGL locus on chromosome 17
The orientation of the goat (Capra hircus) IGL locus on chromosome 17 is reverse (REV).

Legend:
Colors are according to IMGT color menu for genes.
The boxes representing the genes are not to scale. Exons are not shown.
A double slash // indicates a gap in genome assembly. Distances in bp (base pair) or in kb (kilobase) associated with a double slash are taken
into account in the length of the lines and included in the numbers displayed at the right end of the lines.
IGL gene names are according to the IMGT nomenclature [2].
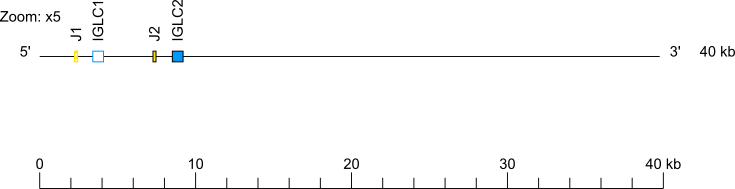
The Capra hircus IGL J-C-CLUSTER comprises 2 cassettes indicated by the numbers 1 and 2 (IGLJ1-IGLC1 and IGLJ2-IGLC2, respectively).
Single arrows show genes whose polarity is opposite to that of the J-C-CLUSTER.
There are two gaps in the locus, one at each end of the inversion, the one in 3', localized in the 5'UTR of IGLV1-37 (complement(236641..236646)) is not represented.
Gene names of IGLV pseudogenes with frameshift(s) in the V-REGION are not displayed due to a lack of space in the locus map at the IMGT standardized scale (line = 200 kb). For all gene names, see Zoom x2.
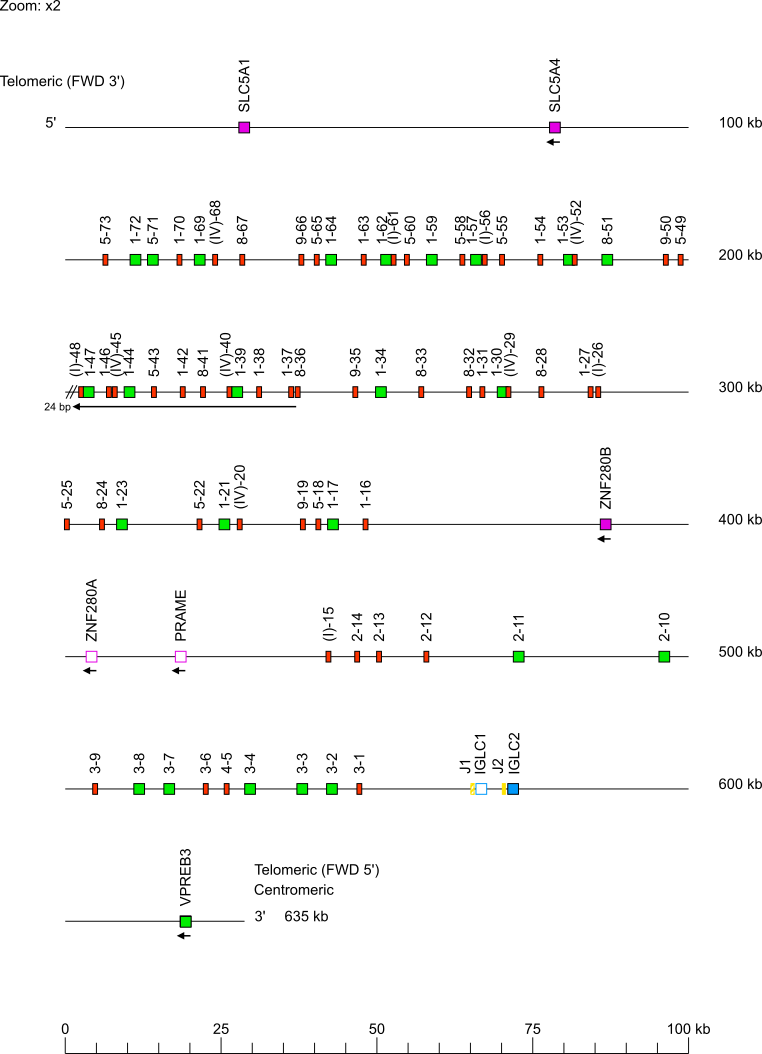
Zoom for IGL V-J-C-CLUSTER

Zoom for IGL J-C-CLUSTER
- [1] Schwartz J.C. et al. Immunogenetics 2018 May;70(5):317-326. doi: 10.1007/s00251-017-1033-3. Epub 2017 Oct 23. Free PMC Article. PMID:29063126
- [2] Lefranc M-P. Front Immunol. 2014 Feb 05;5:22. doi: 10.3389/fimmu.2014.00022. Open access. PMID:24600447
- Created:
- 27/06/2019
- Last updated:
- 22/09/2023
- Authors:
- Morgane Bertignac
