IMGT Repertoire (IG and TR)
Locus representation: horse (Equus caballus) IGH locus on chromosome 24
Horse (Equus caballus) IGH locus on chromosome 24
The orientation of the horse (Equus caballus) IGH locus on chromosome 24 is reverse (REV).
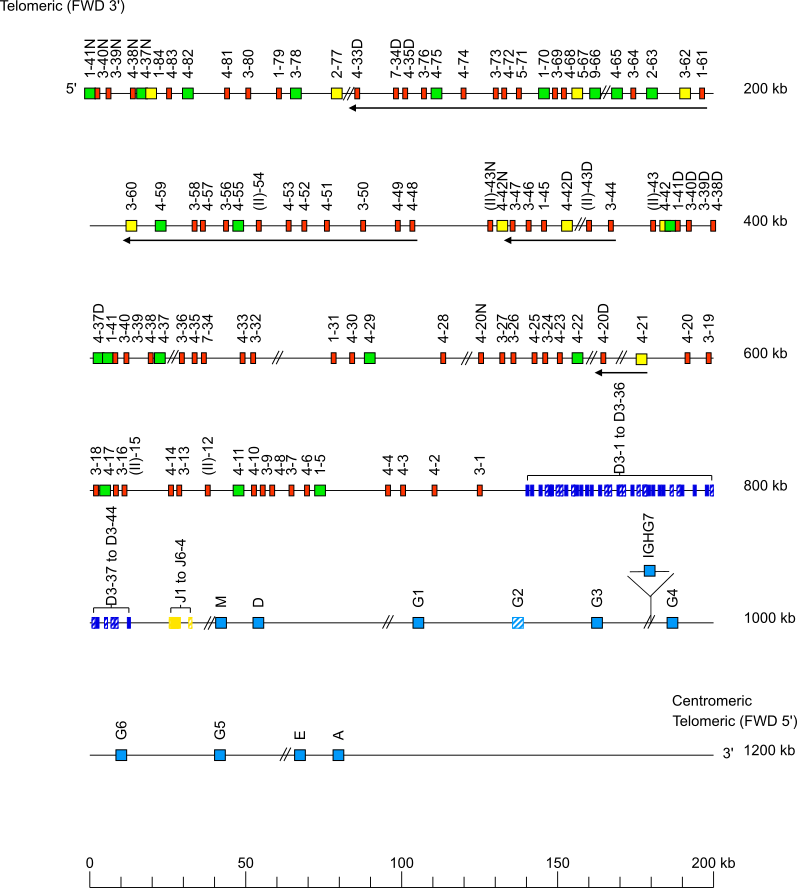
Legend:
Colors are according to IMGT color menu for genes.
The boxes representing the genes are not to scale. Exons are not shown.
A double slash // indicates a gap in genome assembly. Distances in bp (base pair) or in kb (kilobase) associated with a double slash are taken
into account in the length of the lines and included in the numbers displayed at the right end of the lines.
IGH gene names are according to the IMGT nomenclature [1].
Single arrows show genes whose polarity is opposite to that of the D-J-C-CLUSTER comprising IGHD (D1 to D40)-IGHJ (J1 to J9)-IGHC (IGHM-IGHD-IGHG1-IGHG2-IGHG3-IGHG4-IGHG6-IGHG5-IGHE-IGHA).
There are several gaps in the locus, the follow ones are not represented: the gap between IGHV4-37D and IGHV1-41, the gap between IGHV4-42 and GHV1-41D and the one between IGHV4-37D and IGHV1-41.
The IGHG7 constant gene is not found in the EquCab3.0 assembly, there is a gap at this position (IMGT000040 accession number). However, IGHG7 is found in the accession numbers NW_019645282 and in AY445517 [2]
Zoom for IGHD and IGHJ genes

- [1] Lefranc M-P. Front Immunol. 2014 Feb 05;5:22. doi: 10.3389/fimmu.2014.00022. Open access. PMID:24600447
- [2] Wagner B. et al., J. Immunol. 2004 Sep 1;173(5):3230-42. doi: 10.4049/jimmunol.173.5.3230.. PMID:15322185
- Created:
- 09/06/2020
- Last updated:
- 22/09/2023
- Authors:
- Joumana Jabado-Michaloud, Marie-Paule Lefranc and Sofia Kossida
